What is Promethease?
Promethease is an online tool that analyzes raw DNA files inputted by users to generate personalized DNA reports based on genetic literature.
The information in the report is based on SNPedia, a wiki-based online database that summarizes health and ancestral implications of genetic variants.
Though popular, it is often criticized to be too technical and difficult to read. You can take a look at Promethease's sample report.
Update: In 2019, MyHeritage acquired Promethease and SNPedia
MyHeritage offered Promethease free of charge through the end of 2019 and continues to maintain SNPedia as a free resource for academic and non-profit users. For non-European users, the raw DNA data will be shifted to MyHeritage and added to new accounts, which will be created for them. However, the users will retain the ownership of their DNA raw data file and are free to delete it from MyHeritage’s server.
How Does Promethease Work?
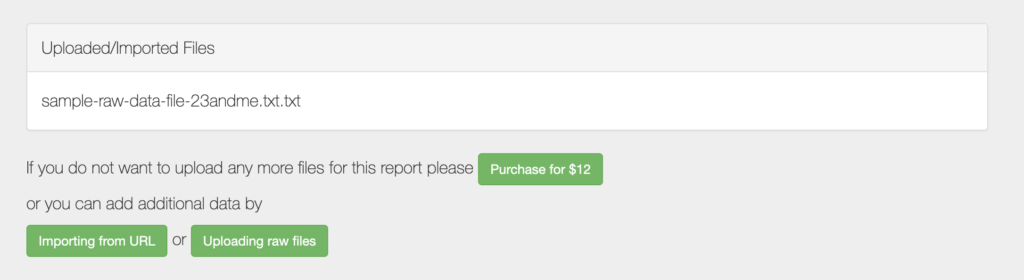
- DNA report generation: For a fee of $12, users can upload their raw DNA data files to receive a comprehensive report that details the genetic variants they carry and their associations with health conditions, traits, and drug responses. Reports are generated in approximately 10-20 minutes after submission.
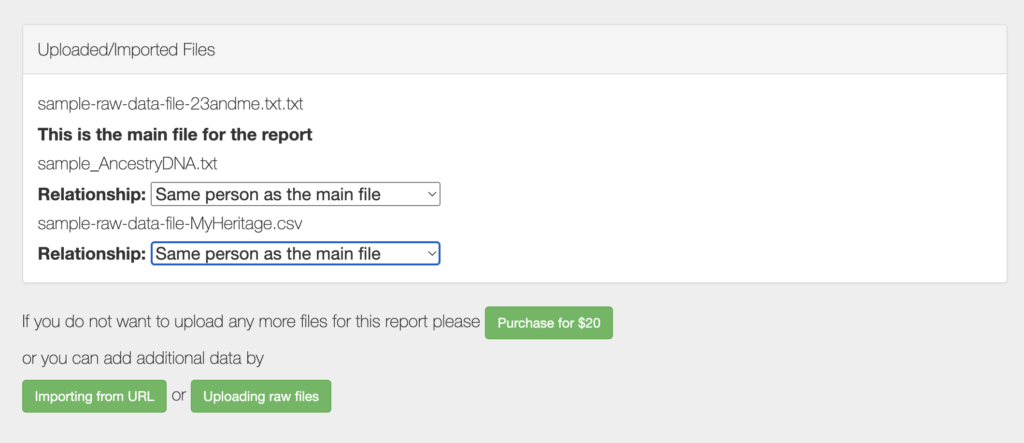
- Multiple file uploads: For an additional charge of $4 per file, users can combine multiple raw data files into one report. This feature is useful for comparing genetic data from different testing services or analyzing the genetic data of multiple individuals.
- Access to scientific literature: The reports link genetic variants to relevant scientific studies, allowing users to explore the implications of their genetic data in depth. This aspect is particularly valuable for those interested in understanding the scientific context of their genetic information.
What Is Included In a Promethease Report?
A Promethease report provides a comprehensive analysis of your genetic data, covering various aspects such as:
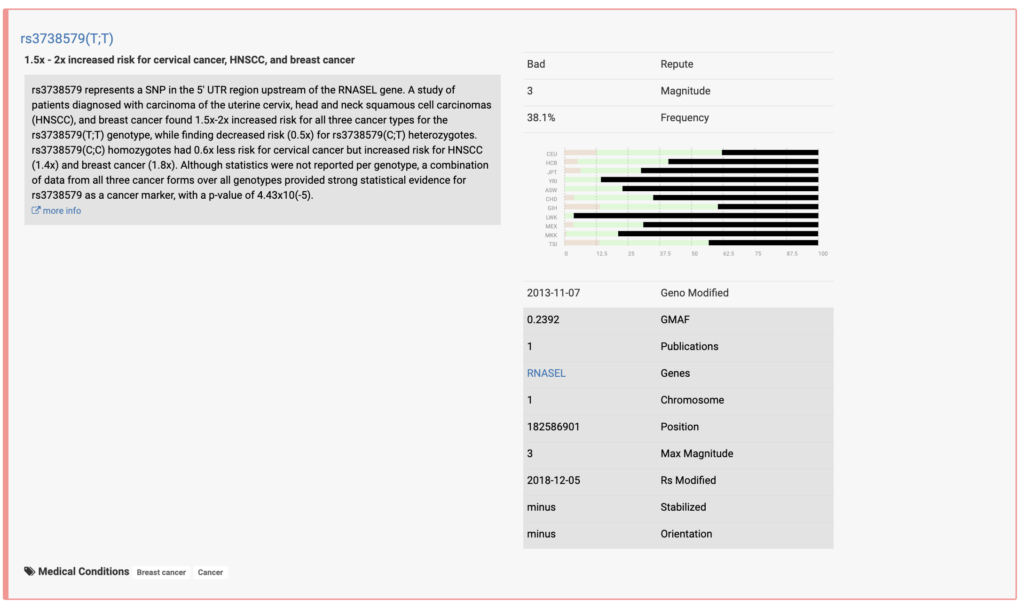
- Health Conditions: The report highlights genetic variants associated with increased or decreased risk for certain health conditions like heart disease, cancer, Alzheimer's, etc.
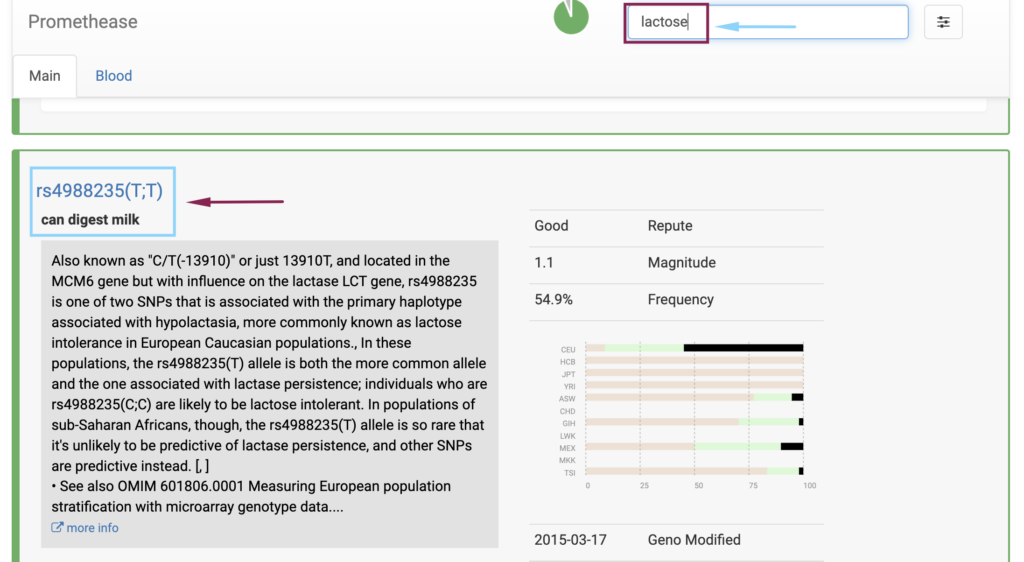
- Traits: It includes information on genetic factors influencing physical traits, such as eye color, hair color, lactose intolerance, and more.
- Medication Response: The report indicates how your genetic makeup may impact your response to certain medications, helping guide personalized treatment decisions.
- Ancestry: While not a primary focus, Promethease can provide some insights into your ancestral background based on your genetic data.
How to Filter and Search the Report
- Promethease uses a wiki-style format, allowing you to search for specific genes, variants, or conditions of interest, similar to how you would use Wikipedia.
- The report is divided into sections for different categories like health, traits, medications, etc. You can navigate to the relevant section to find information on a particular topic.
- Each genetic variant is assigned a "magnitude score," indicating its importance. You can focus on variants with higher magnitude scores that are more likely to have a significant impact.
- The "Summary" column provides a concise interpretation of how your genotype relates to the associated trait or condition. Look for terms like "increased risk," "decreased risk," or "typical."
Understanding The Promethease Report
Before you get into reading your Promethease report, it will help to know the meaning of a few terms that are frequently used in them. They are tabulated below.
| Term | Description |
|---|---|
| Repute | This is a parameter that is applied to a genotype. It can be good, bad or not set. This is based on a general opinion. |
| Magnitude | It tells you how interesting your genotype is. It is purely a subjective parameter. It is a numerical value between 0 and 10, with 0 assigned to boring information and 10 being assigned to really significant information. This can be set to whichever value you prefer. Most users set this value up to a moderate 5. |
| Orientation | As we know, genetic studies are based on genes. Orientation is simply the chromosome position of these genes. These positions are based on a reference human genome. The present build is GRCh38 patch 12, that was released in December 2017 (you don’t have to remember this!). |
| Frequency | How rare or common a genotype is in your population. |
| Geno Time | Timestamp when the genotype page was last updated. This is with reference to SNPedia. |
| ClinVar | If you select that button, only health conditions that are found in ClinVar will be displayed. ClinVar is a public archive of reports of the relationships among human gene variations and phenotypes, with supporting evidence. |
| Genosets (gs IDs) | A genoset is a defined set of genotypes that are associated with a particular phenotype. |
| SNPs for the other sex | Many people are taken aback by this term. Don’t worry about this. It simply tells you that manifestations of these SNPs are irrelevant to your sex. For example it is irrelevant to a woman, how an SNP can increase the risk for prostate cancer. |
| GMAF | Stands for Global Minor Allele Frequency. The allele (an alternative form of your gene) that is less common in the global gene pool. This is as per the 1000 Genomes Project. |
| Carrier | Many of you may already be familiar with this term. Carriers are individuals who carry only one copy the risky allele. It is of significance only in disorders that are know to be caused due to a single gene mutation like Cystic Fibrosis or Huntington’s disease (a.k.a Mendelian disorders). Carriers do not manifest the disease but may pass on the mutant allele to the next generation. |
| CI | Stands for Confidence Interval. The range within which a value can be deemed true with a certain degree of confidence. It is usually taken as 95%. |
| de novo mutation | True to the name, it means a new mutation that was not found in both your parents. |
| CNV | Stands for Copy Number Variants. Number of copies of the gene you have in your genome. Large copy number of genes are commonly found in many forms of cancer. |
| Haplotype | A specific combination of SNPs all occurring on the same chromosome. This is all chromosomes inherited from your mother or father. |
| Diplotype | It refers to a specific combination of haplotypes. |
| Familial risk | It is a comparison between the risk of individuals with relatives with a disease to the risk of individuals with relatives who do not have the same disease. |
| GWAS | Stands for Genome-Wide Association Study. A study of the markers (usually SNPs) across the entire genome to see which ones are statistically more or less common in one group of people (often patients with a specific disease) compared to another group of people (typically people unaffected by that disease). |
| In/dels | They are a type of polymorphism that involves the insertion or deletion of slightly bigger sequence of bases. |
| SNP | Stands for Single Nucleotide Polymorphism. As the name suggests, it is a change occuring in only one base. |
| Odds ratio (OR) | It is a useful number to evaluate the association between an allele or genotype with a phenotype. The phenotype might be an inherited trait (like widow’s peak) or a disease (like Type 2 Diabetes). This number is arrived at by comparing carriers of a less common allele to people with 2 copies of the most common allele. |
| Penetrance | The degree to which you're likely to actually have a given trait or phenotype when you carry a given variation. Penetrance is considered high if over 80%, and moderate if between 20 - 80%. |
| Position | Location of an SNP on the chromosome. This position is however only relative to positions in the reference genome sequence. You cannot assign absolute positions to SNPs on a chromosome. |
| Prevalence | The proportion of people with a given condition at a given time. Usually a percentage or a ratio. |
| rs# | All SNPs have an rs identifier that is officially assigned by dbSNP, a public database of SNPs maintained by the National Institute of Health (NIH). |
| Wild-type | The genotype or the allele that is most common in the population and serves as a reference to evaluate all other types of alleles. |
Limitations Of Promethease Reports
Too Much Information
A Promthease genetic report typically contains around 25,000 entries which can be very overwhelming, especially for those without a genetic background.
However, as per the sample files they have for analysis, only 24% of these entries are considered "good."
Complex Terminology
The reports frequently use scientific and technical language, making them difficult to understand for laypersons.
This complexity can lead to confusion and misinterpretation of the results
No Personalized Guidance/Recommendations
Promethease reports primarily present data without personalized interpretation or actionable next steps.
There are other genetic analysis services like Xcode Life, which offer tailored advice or recommendations based on the findings.
Poor User Interface
Users have criticized the interface for being user-unfriendly, which can make navigation and understanding the report more challenging.
Some users have expressed frustration with the overall experience of using the platform.
Further, the reports are primarily text-based and lack visual aids that could help users better comprehend their genetic information.
Risk Of False Positives
Some reviews suggest that Promethease analysis is known to produce false positives for conditions like breast cancer.
Not A Diagnostic Tool
Promethease provides risk estimates based on genetic variants but is not intended for medical diagnosis.
There's no obvious mention of this in their reports.
Thus, there's a risk of users misinterpreting the information, leading to unnecessary anxiety or inappropriate health decisions.
Check out this page for more sites that interpret your DNA raw data info
Xcode Life vs. Promethease Raw DNA Analysis
| Xcode Life | Promethease | |
| No. of report categories | 11 | 4 |
| Easy-to-understand reports? | Yes | No |
| Actionable insights | Included | Not included |
| Information sources | Several reputable journals and databases like JAMA, ClinVar, etc. | Only SNPedia |
| Personalized recommendations | Included | Not included |
| Sample reports | Present | Present |
| Report interpretation assistance | Provided | Not provided |




I am still confused by even the simplified report. In one area, it says bad for a certain health issue, but then for the same health issue, there are also "good" reports. I wish there was a summary that says, "based on your, whatever (weighing out the good and bad) you may be at higher risk for _______. Even though you have good ____________. " LOL! Seriously, I am not any less confused by the simplified report than I was by the original
Hello! Please drop an email with this query to hello+support@xcode.life; one of the report developers will get in touch with you. Hope this clarifies.
I have tried Promethease, Genetic Genie and Self Decode. I am still looking for a good raw data service. Is Xcode Life really as good as the reviews?
Great analysis! Good insights into how I can use my raw data results. Thank you for this.
Has prometheus hiked their price?
Thank you for this article. Promethease sure confused me. I think ill go and have a look at the report again.
Will the Simplified Promethease report give me all the information from my Promethease report?
Great! Thanks! I almost gave up trying to understand my Promethease report. Will go back and take a look now after reading this. Also the Simplified Promethease report looks interesting. Does it use the Promethease report or the genome raw data file?
Hi Cathy. Thank you for stopping by. To get the Simplified Promethease report you will have to upload your Promethease report from 2017 or later.
Where can I order the Simplified Promethease report?
Hi Mike. You can place an order for the Simplified Promethease report from here: https://www.xcode.life/product/ftdna-ancestrydna-23andme-raw-data-interpretation-analysis-tools
Choose Promethease as your service provider and the Simplified Promethease report will show up as the product.
Wow! I thought I was the only one who was confused by my Promethease report. I am getting the Simplified Promethease report right away!
Wow! Thanks for this. I finally know have some idea of what is there in my Promethease report.