Haplogroup K
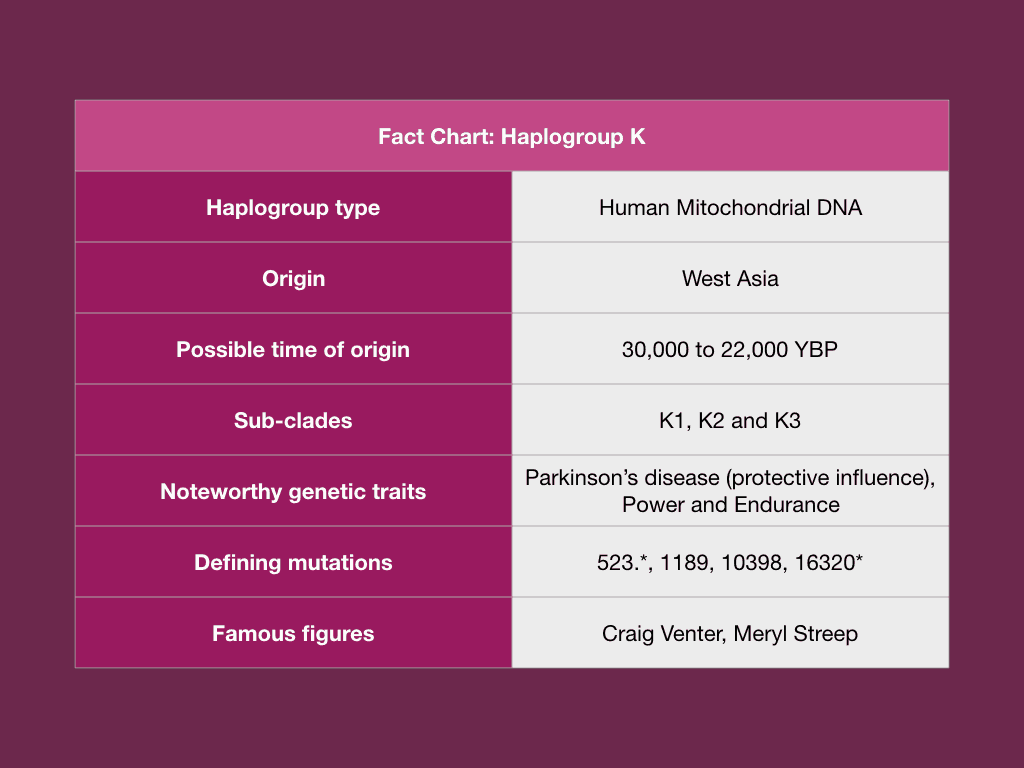
Haplogroup K is a mitochondrial haplogroup (mtDNA haplogroup) that is estimated to have originated during the Upper Paleolithic or Late Stone Age around 30,000 to 22,000 ybp. The most common sub-clade of this haplogroup is U8b. According to a study conducted in a Caucasian population, individuals who were found to be classifed under haplogroup K showed a significantly lower risk toward Parkinson’s disease as compared to individuals assigned the haplogroup H.
Important! Remember to check your raw data for Y-DNA (if you are male) and mt-DNA markers. Your autosomal DNA raw data is not enough for haplogroup analysis.
Origin and Distribution
The geographical distribution of Haplogroup K ranges from Northwest Europe to regions of the Eastern Mediterranean and the Middle East. The highest frequencies of Haplogroup K was found to be in Belgium, The Netherlands, Cyprus and Lebanon.
Haplogroup K was found to have originated in West Asia between 20,000 and 38,000 ybp as a sub-clade of U8b. Sub-clades of haplogroup have their origins in different regions. For example, the sub-clades K1a, K1b and K2a were discovered among the Early Neolithic farmers from the Near East. At the same time sub-clades K1c, K2b and K2c were found among the Neolithic farmers who do not have any Near Eastern roots. Today, this haplogroup is most common among eastern Europe and is postulated to have spread to regions of Germany and Central Asia during the Bronze Age along with Y-Haplogroup R1a.
Interesting trivia: According to the book The Seven Daughters of Eve by Bryan Sykes, the originator of Haplogroup K was given the name Katrine.
History
Haplogroup K is the most common haplogroup among the Ashkenazi Jewish population. The highest percentage of lineages from this population that belong to haplogroup K. Using ancient mtDNA samples of Neolithic farmers, it was found that around, around 15% of these samples belong to haplogroup K. As these farmers mingled with the aboriginal Europeans descended from Mesolithic fishermen and hunter-gatherers the frequency of this haplogroup decreased.
The K1a sub-clade of this haplogroup is estimated to have originated 19,000 to 22,000 ybp. Among the K haplogroup sub-clades, this is the most common in Europe. This includes regions of Spain, Catalonia, Portugal, Hungary and Sweden. K1a is also predominant in the Levant mostly among the Druzes. The Druze population is known to be the closest population to the original Neolithic farmers. They are also said to be constituting the early populations of the Levant before the Arabic expansion. The Druze population have 13% frequency of haplogroup K with sub-clade K1a being the most prevalent.
Get your Origin 2.0 Ancestry report and find out your Haplogroup!
Genetic influence
Parkinson’s Disease
A 2009 study by Rollins et al, showed an association between mitochondrial DNA (mtDNA) alleles and the brain. The mutation A10398G which defines K1 was found to be associated with increased pH. An increased brain pH was found to protect individuals against Parkinson’s disease, schizophrenia, bipolar disorder and depression.
Power and Endurance
Studies conducted on elite athletes from the Finnish population showed that haplogroup K was not found among any of the endurance athletes (Niemi and Majamaa, 2005). Another study involving 395 elite Polish athletes- 213 endurance and 182 power and 413 sedentary individuals as control demonstrated an underrepresentation of haplogroup K among the athletes as compared to the controls.